Remote controlling#
DataLab may be controlled remotely using the XML-RPC protocol which is natively supported by Python (and many other languages). Remote controlling allows to access DataLab main features from a separate process.
Note
If you are looking for a lighweight alternative solution to remote control DataLab (i.e. without having to install the whole DataLab package and its dependencies on your environment), please have a look at the DataLab Simple Client package (pip install cdlclient).
From an IDE#
DataLab may be controlled remotely from an IDE (e.g. Spyder or any other
IDE, or even a Jupyter Notebook) that runs a Python script. It allows to
connect to a running DataLab instance, adds a signal and an image, and then
runs calculations. This feature is exposed by the RemoteProxy class that
is provided in module cdl.proxy.
From a third-party application#
DataLab may also be controlled remotely from a third-party application, for the same purpose.
If the third-party application is written in Python 3, it may directly use the RemoteProxy class as mentioned above. From another language, it is also achievable, but it requires to implement a XML-RPC client in this language using the same methods of proxy server as in the RemoteProxy class.
Data (signals and images) may also be exchanged between DataLab and the remote client application, in both directions.
The remote client application may be written in any language that supports XML-RPC. For example, it is possible to write a remote client application in Python, Java, C++, C#, etc. The remote client application may be a graphical application or a command line application.
The remote client application may be run on the same computer as DataLab or on a different computer. In the latter case, the remote client application must know the IP address of the computer running DataLab.
The remote client application may be run before or after DataLab. In the latter case, the remote client application must try to connect to DataLab until it succeeds.
Supported features#
Supported features are the following:
Switch to signal or image panel
Remove all signals and images
Save current session to a HDF5 file
Open HDF5 files into current session
Browse HDF5 file
Open a signal or an image from file
Add a signal
Add an image
Get object list
Run calculation with parameters
Note
The signal and image objects are described on this section: Internal data model.
Some examples are provided to help implementing such a communication between your application and DataLab:
See module:
cdl.tests.remoteclient_appSee module:
cdl.tests.remoteclient_unit
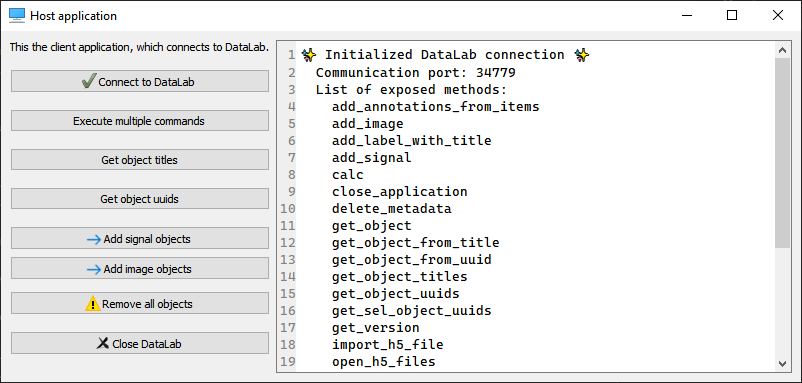
Screenshot of remote client application test (cdl.tests.remoteclient_app)#
Examples#
When using Python 3, you may directly use the RemoteProxy class as in examples cited above or below.
Here is an example in Python 3 of a script that connects to a running DataLab instance, adds a signal and an image, and then runs calculations (the cell structure of the script make it convenient to be used in Spyder IDE):
"""
Example of remote control of DataLab current session,
from a Python script running outside DataLab (e.g. in Spyder)
Created on Fri May 12 12:28:56 2023
@author: p.raybaut
"""
# %% Importing necessary modules
# NumPy for numerical array computations:
import numpy as np
# DataLab remote control client:
from cdlclient import SimpleRemoteProxy as RemoteProxy
# %% Connecting to DataLab current session
proxy = RemoteProxy()
# %% Executing commands in DataLab (...)
z = np.random.rand(20, 20)
proxy.add_image("toto", z)
# %% Executing commands in DataLab (...)
proxy.toggle_auto_refresh(False) # Turning off auto-refresh
x = np.array([1.0, 2.0, 3.0])
y = np.array([4.0, 5.0, -1.0])
proxy.add_signal("toto", x, y)
# %% Executing commands in DataLab (...)
proxy.compute_derivative()
proxy.toggle_auto_refresh(True) # Turning on auto-refresh
# %% Executing commands in DataLab (...)
proxy.set_current_panel("image")
# %% Executing a lot of commands without refreshing DataLab
z = np.random.rand(400, 400)
proxy.add_image("foobar", z)
with proxy.context_no_refresh():
for _idx in range(100):
proxy.compute_fft()
Here is a Python 2.7 reimplementation of this class:
# Copyright (c) DataLab Platform Developers, BSD 3-Clause license, see LICENSE file.
"""
DataLab remote controlling class for Python 2.7
"""
import io
import os
import os.path as osp
import socket
import sys
import ConfigParser as cp
import numpy as np
from guidata.userconfig import get_config_dir
from xmlrpclib import Binary, ServerProxy
def array_to_rpcbinary(data):
"""Convert NumPy array to XML-RPC Binary object, with shape and dtype"""
dbytes = io.BytesIO()
np.save(dbytes, data, allow_pickle=False)
return Binary(dbytes.getvalue())
def get_cdl_xmlrpc_port():
"""Return DataLab current XML-RPC port"""
if sys.platform == "win32" and "HOME" in os.environ:
os.environ.pop("HOME") # Avoid getting old WinPython settings dir
fname = osp.join(get_config_dir(), ".DataLab", "DataLab.ini")
ini = cp.ConfigParser()
ini.read(fname)
try:
return ini.get("main", "rpc_server_port")
except (cp.NoSectionError, cp.NoOptionError):
raise ConnectionRefusedError("DataLab has not yet been executed")
class RemoteClient(object):
"""Object representing a proxy/client to DataLab XML-RPC server"""
def __init__(self):
self.port = None
self.serverproxy = None
def connect(self, port=None):
"""Connect to DataLab XML-RPC server"""
if port is None:
port = get_cdl_xmlrpc_port()
self.port = port
url = "http://127.0.0.1:" + port
self.serverproxy = ServerProxy(url, allow_none=True)
try:
self.get_version()
except socket.error:
raise ConnectionRefusedError("DataLab is currently not running")
def get_version(self):
"""Return DataLab public version"""
return self.serverproxy.get_version()
def close_application(self):
"""Close DataLab application"""
self.serverproxy.close_application()
def raise_window(self):
"""Raise DataLab window"""
self.serverproxy.raise_window()
def get_current_panel(self):
"""Return current panel"""
return self.serverproxy.get_current_panel()
def set_current_panel(self, panel):
"""Switch to panel"""
self.serverproxy.set_current_panel(panel)
def reset_all(self):
"""Reset all application data"""
self.serverproxy.reset_all()
def toggle_auto_refresh(self, state):
"""Toggle auto refresh state"""
self.serverproxy.toggle_auto_refresh(state)
def toggle_show_titles(self, state):
"""Toggle show titles state"""
self.serverproxy.toggle_show_titles(state)
def save_to_h5_file(self, filename):
"""Save to a DataLab HDF5 file"""
self.serverproxy.save_to_h5_file(filename)
def open_h5_files(self, h5files, import_all, reset_all):
"""Open a DataLab HDF5 file or import from any other HDF5 file"""
self.serverproxy.open_h5_files(h5files, import_all, reset_all)
def import_h5_file(self, filename, reset_all):
"""Open DataLab HDF5 browser to Import HDF5 file"""
self.serverproxy.import_h5_file(filename, reset_all)
def load_from_files(self, filenames):
"""Open objects from files in current panel (signals/images)"""
self.serverproxy.load_from_files(filenames)
def add_signal(
self, title, xdata, ydata, xunit=None, yunit=None, xlabel=None, ylabel=None
):
"""Add signal data to DataLab"""
xbinary = array_to_rpcbinary(xdata)
ybinary = array_to_rpcbinary(ydata)
p = self.serverproxy
return p.add_signal(title, xbinary, ybinary, xunit, yunit, xlabel, ylabel)
def add_image(
self,
title,
data,
xunit=None,
yunit=None,
zunit=None,
xlabel=None,
ylabel=None,
zlabel=None,
):
"""Add image data to DataLab"""
zbinary = array_to_rpcbinary(data)
p = self.serverproxy
return p.add_image(title, zbinary, xunit, yunit, zunit, xlabel, ylabel, zlabel)
def get_object_titles(self, panel=None):
"""Get object (signal/image) list for current panel"""
return self.serverproxy.get_object_titles(panel)
def get_object(self, nb_id_title=None, panel=None):
"""Get object (signal/image) by number, id or title"""
return self.serverproxy.get_object(nb_id_title, panel)
def get_object_uuids(self, panel=None, group=None):
"""Get object (signal/image) list for current panel"""
return self.serverproxy.get_object_uuids(panel, group)
def test_remote_client():
"""DataLab Remote Client test"""
cdl = RemoteClient()
cdl.connect()
data = np.array([[3, 4, 5], [7, 8, 0]], dtype=np.uint16)
cdl.add_image("toto", data)
if __name__ == "__main__":
test_remote_client()
Connection dialog#
The DataLab package also provides a connection dialog that may be used
to connect to a running DataLab instance. It is exposed by the
cdl.widgets.connection.ConnectionDialog class.
Screenshot of connection dialog (cdl.widgets.connection.ConnectionDialog)#
Example of use:
# Copyright (c) DataLab Platform Developers, BSD 3-Clause license, see LICENSE file.
"""
DataLab Remote client connection dialog example
"""
# guitest: show,skip
from guidata.qthelpers import qt_app_context
from qtpy import QtWidgets as QW
from cdl.proxy import RemoteProxy
from cdl.widgets.connection import ConnectionDialog
def test_dialog():
"""Test connection dialog"""
proxy = RemoteProxy(autoconnect=False)
with qt_app_context():
dlg = ConnectionDialog(proxy.connect)
if dlg.exec():
QW.QMessageBox.information(None, "Connection", "Successfully connected")
else:
QW.QMessageBox.critical(None, "Connection", "Connection failed")
if __name__ == "__main__":
test_dialog()
Public API: remote client#
- class cdl.core.remote.RemoteClient[source]#
Object representing a proxy/client to DataLab XML-RPC server. This object is used to call DataLab functions from a Python script.
Examples
Here is a simple example of how to use RemoteClient in a Python script or in a Jupyter notebook:
>>> from cdl.core.remote import RemoteClient >>> proxy = RemoteClient() >>> proxy.connect() Connecting to DataLab XML-RPC server...OK (port: 28867) >>> proxy.get_version() '1.0.0' >>> proxy.add_signal("toto", np.array([1., 2., 3.]), np.array([4., 5., -1.])) True >>> proxy.get_object_titles() ['toto'] >>> proxy["toto"] <cdl.core.model.signal.SignalObj at 0x7f7f1c0b4a90> >>> proxy[1] <cdl.core.model.signal.SignalObj at 0x7f7f1c0b4a90> >>> proxy[1].data array([1., 2., 3.])
- connect(port: str | None = None, timeout: float | None = None, retries: int | None = None) None[source]#
Try to connect to DataLab XML-RPC server.
- Parameters:
port – XML-RPC port to connect to. If not specified, the port is automatically retrieved from DataLab configuration.
timeout – Timeout in seconds. Defaults to 5.0.
retries – Number of retries. Defaults to 10.
- Raises:
ConnectionRefusedError – Unable to connect to DataLab
ValueError – Invalid timeout (must be >= 0.0)
ValueError – Invalid number of retries (must be >= 1)
- add_signal(title: str, xdata: ndarray, ydata: ndarray, xunit: str | None = None, yunit: str | None = None, xlabel: str | None = None, ylabel: str | None = None) bool[source]#
Add signal data to DataLab.
- Parameters:
title – Signal title
xdata – X data
ydata – Y data
xunit – X unit. Defaults to None.
yunit – Y unit. Defaults to None.
xlabel – X label. Defaults to None.
ylabel – Y label. Defaults to None.
- Returns:
True if signal was added successfully, False otherwise
- Raises:
ValueError – Invalid xdata dtype
ValueError – Invalid ydata dtype
- add_image(title: str, data: ndarray, xunit: str | None = None, yunit: str | None = None, zunit: str | None = None, xlabel: str | None = None, ylabel: str | None = None, zlabel: str | None = None) bool[source]#
Add image data to DataLab.
- Parameters:
title – Image title
data – Image data
xunit – X unit. Defaults to None.
yunit – Y unit. Defaults to None.
zunit – Z unit. Defaults to None.
xlabel – X label. Defaults to None.
ylabel – Y label. Defaults to None.
zlabel – Z label. Defaults to None.
- Returns:
True if image was added successfully, False otherwise
- Raises:
ValueError – Invalid data dtype
- add_object(obj: SignalObj | ImageObj) None[source]#
Add object to DataLab.
- Parameters:
obj – Signal or image object
- calc(name: str, param: DataSet | None = None) None[source]#
Call compute function
namein current panel’s processor.- Parameters:
name – Compute function name
param – Compute function parameter. Defaults to None.
- Raises:
ValueError – unknown function
- get_object(nb_id_title: int | str | None = None, panel: str | None = None) SignalObj | ImageObj[source]#
Get object (signal/image) from index.
- Parameters:
nb_id_title – Object number, or object id, or object title. Defaults to None (current object).
panel – Panel name. Defaults to None (current panel).
- Returns:
Object
- Raises:
KeyError – if object not found
- get_object_shapes(nb_id_title: int | str | None = None, panel: str | None = None) list[source]#
Get plot item shapes associated to object (signal/image).
- Parameters:
nb_id_title – Object number, or object id, or object title. Defaults to None (current object).
panel – Panel name. Defaults to None (current panel).
- Returns:
List of plot item shapes
- add_annotations_from_items(items: list, refresh_plot: bool = True, panel: str | None = None) None[source]#
Add object annotations (annotation plot items).
- Parameters:
items – annotation plot items
refresh_plot – refresh plot. Defaults to True.
panel – panel name (valid values: “signal”, “image”). If None, current panel is used.
- add_group(title: str, panel: str | None = None, select: bool = False) None#
Add group to DataLab.
- Parameters:
title – Group title
panel – Panel name (valid values: “signal”, “image”). Defaults to None.
select – Select the group after creation. Defaults to False.
- add_label_with_title(title: str | None = None, panel: str | None = None) None#
Add a label with object title on the associated plot
- Parameters:
title – Label title. Defaults to None. If None, the title is the object title.
panel – panel name (valid values: “signal”, “image”). If None, current panel is used.
- context_no_refresh() Generator[None, None, None]#
Return a context manager to temporarily disable auto refresh.
- Returns:
Context manager
Example
>>> with proxy.context_no_refresh(): ... proxy.add_image("image1", data1) ... proxy.compute_fft() ... proxy.compute_wiener() ... proxy.compute_ifft() ... # Auto refresh is disabled during the above operations
- delete_metadata(refresh_plot: bool = True, keep_roi: bool = False) None#
Delete metadata of selected objects
- Parameters:
refresh_plot – Refresh plot. Defaults to True.
keep_roi – Keep ROI. Defaults to False.
- get_current_panel() str#
Return current panel name.
- Returns:
“signal”, “image”, “macro”))
- Return type:
Panel name (valid values
- get_group_titles_with_object_infos() tuple[list[str], list[list[str]], list[list[str]]]#
Return groups titles and lists of inner objects uuids and titles.
- Returns:
groups titles, lists of inner objects uuids and titles
- Return type:
Tuple
- get_object_titles(panel: str | None = None) list[str]#
Get object (signal/image) list for current panel. Objects are sorted by group number and object index in group.
- Parameters:
panel – panel name (valid values: “signal”, “image”, “macro”). If None, current data panel is used (i.e. signal or image panel).
- Returns:
List of object titles
- Raises:
ValueError – if panel not found
- get_object_uuids(panel: str | None = None, group: int | str | None = None) list[str]#
Get object (signal/image) uuid list for current panel. Objects are sorted by group number and object index in group.
- Parameters:
panel – panel name (valid values: “signal”, “image”). If None, current panel is used.
group – Group number, or group id, or group title. Defaults to None (all groups).
- Returns:
List of object uuids
- Raises:
ValueError – if panel not found
- classmethod get_public_methods() list[str]#
Return all public methods of the class, except itself.
- Returns:
List of public methods
- get_sel_object_uuids(include_groups: bool = False) list[str]#
Return selected objects uuids.
- Parameters:
include_groups – If True, also return objects from selected groups.
- Returns:
List of selected objects uuids.
- import_h5_file(filename: str, reset_all: bool | None = None) None#
Open DataLab HDF5 browser to Import HDF5 file.
- Parameters:
filename – HDF5 file name
reset_all – Reset all application data. Defaults to None.
- load_from_directory(path: str) None#
Open objects from directory in current panel (signals/images).
- Parameters:
path – directory path
- load_from_files(filenames: list[str]) None#
Open objects from files in current panel (signals/images).
- Parameters:
filenames – list of file names
- open_h5_files(h5files: list[str] | None = None, import_all: bool | None = None, reset_all: bool | None = None) None#
Open a DataLab HDF5 file or import from any other HDF5 file.
- Parameters:
h5files – List of HDF5 files to open. Defaults to None.
import_all – Import all objects from HDF5 files. Defaults to None.
reset_all – Reset all application data. Defaults to None.
- run_macro(number_or_title: int | str | None = None) None#
Run macro.
- Parameters:
number_or_title – Macro number, or macro title. Defaults to None (current macro).
- Raises:
ValueError – if macro not found
- save_to_h5_file(filename: str) None#
Save to a DataLab HDF5 file.
- Parameters:
filename – HDF5 file name
- select_groups(selection: list[int | str] | None = None, panel: str | None = None) None#
Select groups in current panel.
- Parameters:
selection – List of group numbers (1 to N), or list of group uuids, or None to select all groups. Defaults to None.
panel – panel name (valid values: “signal”, “image”). If None, current panel is used. Defaults to None.
- select_objects(selection: list[int | str], panel: str | None = None) None#
Select objects in current panel.
- Parameters:
selection – List of object numbers (1 to N) or uuids to select
panel – panel name (valid values: “signal”, “image”). If None, current panel is used. Defaults to None.
- set_current_panel(panel: str) None#
Switch to panel.
- Parameters:
panel – Panel name (valid values: “signal”, “image”, “macro”))
- stop_macro(number_or_title: int | str | None = None) None#
Stop macro.
- Parameters:
number_or_title – Macro number, or macro title. Defaults to None (current macro).
- Raises:
ValueError – if macro not found
Public API: additional methods#
The remote control class methods (either using the proxy or the remote client)
may be completed with additional methods which are dynamically added at
runtime. This mechanism allows to access the methods of the processors of DataLab
(see cdl.core.gui.processor).
Signal processor#
When working with signals, the methods of cdl.core.gui.processor.signal.SignalProcessor
may be accessed.
- class cdl.core.gui.processor.signal.SignalProcessor(panel: SignalPanel | ImagePanel, plotwidget: PlotWidget)[source]
Object handling signal processing: operations, processing, analysis
- compute_sum() None[source]
Compute sum with
cdl.computation.signal.compute_addition()
- compute_addition_constant(param: ConstantParam | None = None) None[source]
Compute sum with a constant with
cdl.computation.signal.compute_addition_constant()
- compute_average() None[source]
Compute average with
cdl.computation.signal.compute_addition()and divide by the number of signals
- compute_product() None[source]
Compute product with
cdl.computation.signal.compute_product()
- compute_product_constant(param: ConstantParam | None = None) None[source]
Compute product with a constant with
cdl.computation.signal.compute_product_constant()
- compute_swap_axes() None[source]
Swap data axes with
cdl.computation.signal.compute_swap_axes()
- compute_abs() None[source]
Compute absolute value with
cdl.computation.signal.compute_abs()
- compute_re() None[source]
Compute real part with
cdl.computation.signal.compute_re()
- compute_im() None[source]
Compute imaginary part with
cdl.computation.signal.compute_im()
- compute_astype(param: DataTypeSParam | None = None) None[source]
Convert data type with
cdl.computation.signal.compute_astype()
- compute_log10() None[source]
Compute Log10 with
cdl.computation.signal.compute_log10()
- compute_exp() None[source]
Compute Log10 with
cdl.computation.signal.compute_exp()
- compute_sqrt() None[source]
Compute square root with
cdl.computation.signal.compute_sqrt()
- compute_power(param: PowerParam | None = None) None[source]
Compute power with
cdl.computation.signal.compute_power()
- compute_arithmetic(obj2: SignalObj | None = None, param: ArithmeticParam | None = None) None[source]
Compute arithmetic operation between two signals with
cdl.computation.signal.compute_arithmetic()
- compute_difference(obj2: SignalObj | list[SignalObj] | None = None) None[source]
Compute difference between two signals with
cdl.computation.signal.compute_difference()
- compute_difference_constant(param: ConstantParam | None = None) None[source]
Compute difference with a constant with
cdl.computation.signal.compute_difference_constant()
- compute_quadratic_difference(obj2: SignalObj | list[SignalObj] | None = None) None[source]
Compute quadratic difference between two signals with
cdl.computation.signal.compute_quadratic_difference()
- compute_division(obj2: SignalObj | list[SignalObj] | None = None) None[source]
Compute division between two signals with
cdl.computation.signal.compute_division()
- compute_division_constant(param: ConstantParam | None = None) None[source]
Compute division by a constant with
cdl.computation.signal.compute_division_constant()
- compute_peak_detection(param: PeakDetectionParam | None = None) None[source]
Detect peaks from data with
cdl.computation.signal.compute_peak_detection()
- compute_reverse_x() None[source]
Reverse X axis with
cdl.computation.signal.compute_reverse_x()
- compute_cartesian2polar(param: AngleUnitParam | None = None) None[source]
Convert cartesian to polar coordinates with
cdl.computation.signal.compute_cartesian2polar()
- compute_polar2cartesian(param: AngleUnitParam | None = None) None[source]
Convert polar to cartesian coordinates with
cdl.computation.signal.compute_polar2cartesian().
- compute_normalize(param: NormalizeParam | None = None) None[source]
Normalize data with
cdl.computation.signal.compute_normalize()
- compute_derivative() None[source]
Compute derivative with
cdl.computation.signal.compute_derivative()
- compute_integral() None[source]
Compute integral with
cdl.computation.signal.compute_integral()
- compute_calibration(param: XYCalibrateParam | None = None) None[source]
Compute data linear calibration with
cdl.computation.signal.compute_calibration()
- compute_clip(param: ClipParam | None = None) None[source]
Compute maximum data clipping with
cdl.computation.signal.compute_clip()
- compute_offset_correction(param: ROI1DParam | None = None) None[source]
Compute offset correction with
cdl.computation.signal.compute_offset_correction()
- compute_gaussian_filter(param: GaussianParam | None = None) None[source]
Compute gaussian filter with
cdl.computation.signal.compute_gaussian_filter()
- compute_moving_average(param: MovingAverageParam | None = None) None[source]
Compute moving average with
cdl.computation.signal.compute_moving_average()
- compute_moving_median(param: MovingMedianParam | None = None) None[source]
Compute moving median with
cdl.computation.signal.compute_moving_median()
- compute_wiener() None[source]
Compute Wiener filter with
cdl.computation.signal.compute_wiener()
- compute_lowpass(param: LowPassFilterParam | None = None) None[source]
Compute high-pass filter with
cdl.computation.signal.compute_filter()
- compute_highpass(param: HighPassFilterParam | None = None) None[source]
Compute high-pass filter with
cdl.computation.signal.compute_filter()
- compute_bandpass(param: BandPassFilterParam | None = None) None[source]
Compute band-pass filter with
cdl.computation.signal.compute_filter()
- compute_bandstop(param: BandStopFilterParam | None = None) None[source]
Compute band-stop filter with
cdl.computation.signal.compute_filter()
- compute_fft(param: FFTParam | None = None) None[source]
Compute FFT with
cdl.computation.signal.compute_fft()
- compute_ifft(param: FFTParam | None = None) None[source]
Compute iFFT with
cdl.computation.signal.compute_ifft()
- compute_magnitude_spectrum(param: SpectrumParam | None = None) None[source]
Compute magnitude spectrum with
cdl.computation.signal.compute_magnitude_spectrum()
- compute_phase_spectrum() None[source]
Compute phase spectrum with
cdl.computation.signal.compute_phase_spectrum()
- compute_psd(param: SpectrumParam | None = None) None[source]
Compute power spectral density with
cdl.computation.signal.compute_psd()
- compute_interpolation(obj2: SignalObj | None = None, param: InterpolationParam | None = None)[source]
Compute interpolation with
cdl.computation.signal.compute_interpolation()
- compute_resampling(param: ResamplingParam | None = None)[source]
Compute resampling with
cdl.computation.signal.compute_resampling()
- compute_detrending(param: DetrendingParam | None = None)[source]
Compute detrending with
cdl.computation.signal.compute_detrending()
- compute_convolution(obj2: SignalObj | None = None) None[source]
Compute convolution with
cdl.computation.signal.compute_convolution()
- compute_windowing(param: WindowingParam | None = None) None[source]
Compute windowing with
cdl.computation.signal.compute_windowing()
- compute_allan_variance(param: AllanVarianceParam | None = None) None[source]
Compute Allan variance with
cdl.computation.signal.compute_allan_variance()
- compute_allan_deviation(param: AllanVarianceParam | None = None) None[source]
Compute Allan deviation with
cdl.computation.signal.compute_allan_deviation()
- compute_overlapping_allan_variance(param: AllanVarianceParam | None = None) None[source]
Compute overlapping Allan variance with
cdl.computation.signal.compute_overlapping_allan_variance()
- compute_modified_allan_variance(param: AllanVarianceParam | None = None) None[source]
Compute modified Allan variance with
cdl.computation.signal.compute_modified_allan_variance()
- compute_hadamard_variance(param: AllanVarianceParam | None = None) None[source]
Compute Hadamard variance with
cdl.computation.signal.compute_hadamard_variance()
- compute_total_variance(param: AllanVarianceParam | None = None) None[source]
Compute total variance with
cdl.computation.signal.compute_total_variance()
- compute_time_deviation(param: AllanVarianceParam | None = None) None[source]
Compute time deviation with
cdl.computation.signal.compute_time_deviation()
- compute_all_stability(param: AllanVarianceParam | None = None) None[source]
Compute all stability analysis features using the following functions:
- compute_polyfit(param: PolynomialFitParam | None = None) None[source]
Compute polynomial fitting curve
- compute_fit(title: str, fitdlgfunc: Callable) None[source]
Compute fitting curve using an interactive dialog
- Parameters:
title – Title of the dialog
fitdlgfunc – Fitting dialog function
- compute_multigaussianfit() None[source]
Compute multi-Gaussian fitting curve using an interactive dialog
- compute_fwhm(param: FWHMParam | None = None) dict[str, ResultShape][source]
Compute FWHM with
cdl.computation.signal.compute_fwhm()
- compute_fw1e2() dict[str, ResultShape][source]
Compute FW at 1/e² with
cdl.computation.signal.compute_fw1e2()
- compute_stats() dict[str, ResultProperties][source]
Compute data statistics with
cdl.computation.signal.compute_stats()
- compute_histogram(param: HistogramParam | None = None) dict[str, ResultShape][source]
Compute histogram with
cdl.computation.signal.compute_histogram()
- compute_contrast() dict[str, ResultProperties][source]
Compute contrast with
cdl.computation.signal.compute_contrast()
- compute_x_at_minmax() dict[str, ResultProperties][source]
Compute x at min/max with
cdl.computation.signal.compute_x_at_minmax()
- compute_x_at_y(param: FindAbscissaParam | None = None) dict[str, ResultProperties][source]
Compute x at y with
cdl.computation.signal.compute_x_at_y().
- compute_sampling_rate_period() dict[str, ResultProperties][source]
Compute sampling rate and period (mean and std) with
cdl.computation.signal.compute_sampling_rate_period()
- compute_bandwidth_3db() None[source]
Compute bandwidth at -3dB with
cdl.computation.signal.compute_bandwidth_3db()
- compute_dynamic_parameters(param: DynamicParam | None = None) dict[str, ResultProperties][source]
Compute Dynamic Parameters (ENOB, SINAD, THD, SFDR, SNR) with
cdl.computation.signal.compute_dynamic_parameters()
Image processor#
When working with images, the methods of cdl.core.gui.processor.image.ImageProcessor
may be accessed.
- class cdl.core.gui.processor.image.ImageProcessor(panel: SignalPanel | ImagePanel, plotwidget: PlotWidget)[source]
Object handling image processing: operations, processing, analysis
- compute_normalize(param: NormalizeParam | None = None) None[source]
Normalize data with
cdl.computation.image.compute_normalize()
- compute_sum() None[source]
Compute sum with
cdl.computation.image.compute_addition()
- compute_addition_constant(param: ConstantParam | None = None) None[source]
Compute sum with a constant using
cdl.computation.image.compute_addition_constant()
- compute_average() None[source]
Compute average with
cdl.computation.image.compute_addition()and dividing by the number of images
- compute_product() None[source]
Compute product with
cdl.computation.image.compute_product()
- compute_product_constant(param: ConstantParam | None = None) None[source]
Compute product with a constant using
cdl.computation.image.compute_product_constant()
- compute_logp1(param: LogP1Param | None = None) None[source]
Compute base 10 logarithm using
cdl.computation.image.compute_logp1()
- compute_rotate(param: RotateParam | None = None) None[source]
Rotate data arbitrarily using
cdl.computation.image.compute_rotate()
- compute_rotate90() None[source]
Rotate data 90° with
cdl.computation.image.compute_rotate90()
- compute_rotate270() None[source]
Rotate data 270° with
cdl.computation.image.compute_rotate270()
- compute_fliph() None[source]
Flip data horizontally using
cdl.computation.image.compute_fliph()
- compute_flipv() None[source]
Flip data vertically with
cdl.computation.image.compute_flipv()
- compute_resize(param: ResizeParam | None = None) None[source]
Resize image with
cdl.computation.image.compute_resize()
- compute_binning(param: BinningParam | None = None) None[source]
Binning image with
cdl.computation.image.compute_binning()
- compute_line_profile(param: LineProfileParam | None = None) None[source]
Compute profile along a vertical or horizontal line with
cdl.computation.image.compute_line_profile()
- compute_segment_profile(param: SegmentProfileParam | None = None)[source]
Compute profile along a segment with
cdl.computation.image.compute_segment_profile()
- compute_average_profile(param: AverageProfileParam | None = None) None[source]
Compute average profile with
cdl.computation.image.compute_average_profile()
- compute_radial_profile(param: RadialProfileParam | None = None) None[source]
Compute radial profile with
cdl.computation.image.compute_radial_profile()
- compute_histogram(param: HistogramParam | None = None) None[source]
Compute histogram with
cdl.computation.image.compute_histogram()
- compute_swap_axes() None[source]
Swap data axes with
cdl.computation.image.compute_swap_axes().
- compute_abs() None[source]
Compute absolute value with
cdl.computation.image.compute_abs()
- compute_re() None[source]
Compute real part with
cdl.computation.image.compute_re()
- compute_im() None[source]
Compute imaginary part with
cdl.computation.image.compute_im()
- compute_astype(param: DataTypeIParam | None = None) None[source]
Convert data type with
cdl.computation.image.compute_astype()
- compute_log10() None[source]
Compute Log10 with
cdl.computation.image.compute_log10()
- compute_exp() None[source]
Compute Log10 with
cdl.computation.image.compute_exp()
- compute_arithmetic(obj2: ImageObj | None = None, param: ArithmeticParam | None = None) None[source]
Compute arithmetic operation between two images with
cdl.computation.image.compute_arithmetic()
- compute_difference(obj2: ImageObj | list[ImageObj] | None = None) None[source]
Compute difference between two images with
cdl.computation.image.compute_difference()
- compute_difference_constant(param: ConstantParam | None = None) None[source]
Compute difference with a constant with
cdl.computation.image.compute_difference_constant()
- compute_quadratic_difference(obj2: ImageObj | list[ImageObj] | None = None) None[source]
Compute quadratic difference between two images with
cdl.computation.image.compute_quadratic_difference()
- compute_division(obj2: ImageObj | list[ImageObj] | None = None) None[source]
Compute division between two images with
cdl.computation.image.compute_division()
- compute_division_constant(param: ConstantParam | None = None) None[source]
Compute division by a constant with
cdl.computation.image.compute_division_constant()
- compute_flatfield(obj2: ImageObj | None = None, param: FlatFieldParam | None = None) None[source]
Compute flat field correction with
cdl.computation.image.compute_flatfield()
- compute_calibration(param: ZCalibrateParam | None = None) None[source]
Compute data linear calibration with
cdl.computation.image.compute_calibration()
- compute_clip(param: ClipParam | None = None) None[source]
Compute maximum data clipping with
cdl.computation.image.compute_clip()
- compute_offset_correction(param: ROI2DParam | None = None) None[source]
Compute offset correction with
cdl.computation.image.compute_offset_correction()
- compute_gaussian_filter(param: GaussianParam | None = None) None[source]
Compute gaussian filter with
cdl.computation.image.compute_gaussian_filter()
- compute_moving_average(param: MovingAverageParam | None = None) None[source]
Compute moving average with
cdl.computation.image.compute_moving_average()
- compute_moving_median(param: MovingMedianParam | None = None) None[source]
Compute moving median with
cdl.computation.image.compute_moving_median()
- compute_wiener() None[source]
Compute Wiener filter with
cdl.computation.image.compute_wiener()
- compute_fft(param: FFTParam | None = None) None[source]
Compute FFT with
cdl.computation.image.compute_fft()
- compute_ifft(param: FFTParam | None = None) None[source]
Compute iFFT with
cdl.computation.image.compute_ifft()
- compute_magnitude_spectrum(param: SpectrumParam | None = None) None[source]
Compute magnitude spectrum with
cdl.computation.image.compute_magnitude_spectrum()
- compute_phase_spectrum() None[source]
Compute phase spectrum with
cdl.computation.image.compute_phase_spectrum()
- compute_psd(param: SpectrumParam | None = None) None[source]
Compute Power Spectral Density (PSD) with
cdl.computation.image.compute_psd()
- compute_butterworth(param: ButterworthParam | None = None) None[source]
Compute Butterworth filter with
cdl.computation.image.compute_butterworth()
- compute_threshold(param: ThresholdParam | None = None) None[source]
Compute parametric threshold with
cdl.computation.image.threshold.compute_threshold()
- compute_threshold_isodata() None[source]
Compute threshold using Isodata algorithm with
cdl.computation.image.threshold.compute_threshold_isodata()
- compute_threshold_li() None[source]
Compute threshold using Li algorithm with
cdl.computation.image.threshold.compute_threshold_li()
- compute_threshold_mean() None[source]
Compute threshold using Mean algorithm with
cdl.computation.image.threshold.compute_threshold_mean()
- compute_threshold_minimum() None[source]
Compute threshold using Minimum algorithm with
cdl.computation.image.threshold.compute_threshold_minimum()
- compute_threshold_otsu() None[source]
Compute threshold using Otsu algorithm with
cdl.computation.image.threshold.compute_threshold_otsu()
- compute_threshold_triangle() None[source]
Compute threshold using Triangle algorithm with
cdl.computation.image.threshold.compute_threshold_triangle()
- compute_threshold_yen() None[source]
Compute threshold using Yen algorithm with
cdl.computation.image.threshold.compute_threshold_yen()
- compute_all_threshold() None[source]
Compute all threshold algorithms using the following functions:
- compute_adjust_gamma(param: AdjustGammaParam | None = None) None[source]
Compute gamma correction with
cdl.computation.image.exposure.compute_adjust_gamma()
- compute_adjust_log(param: AdjustLogParam | None = None) None[source]
Compute log correction with
cdl.computation.image.exposure.compute_adjust_log()
- compute_adjust_sigmoid(param: AdjustSigmoidParam | None = None) None[source]
Compute sigmoid correction with
cdl.computation.image.exposure.compute_adjust_sigmoid()
- compute_rescale_intensity(param: RescaleIntensityParam | None = None) None[source]
Rescale image intensity levels with :py:func`cdl.computation.image.exposure.compute_rescale_intensity`
- compute_equalize_hist(param: EqualizeHistParam | None = None) None[source]
Histogram equalization with
cdl.computation.image.exposure.compute_equalize_hist()
- compute_equalize_adapthist(param: EqualizeAdaptHistParam | None = None) None[source]
Adaptive histogram equalization with
cdl.computation.image.exposure.compute_equalize_adapthist()
- compute_denoise_tv(param: DenoiseTVParam | None = None) None[source]
Compute Total Variation denoising with
cdl.computation.image.restoration.compute_denoise_tv()
- compute_denoise_bilateral(param: DenoiseBilateralParam | None = None) None[source]
Compute bilateral filter denoising with
cdl.computation.image.restoration.compute_denoise_bilateral()
- compute_denoise_wavelet(param: DenoiseWaveletParam | None = None) None[source]
Compute Wavelet denoising with
cdl.computation.image.restoration.compute_denoise_wavelet()
- compute_denoise_tophat(param: MorphologyParam | None = None) None[source]
Denoise using White Top-Hat with
cdl.computation.image.restoration.compute_denoise_tophat()
- compute_all_denoise(params: list | None = None) None[source]
Compute all denoising filters using the following functions:
- compute_white_tophat(param: MorphologyParam | None = None) None[source]
Compute White Top-Hat with
cdl.computation.image.morphology.compute_white_tophat()
- compute_black_tophat(param: MorphologyParam | None = None) None[source]
Compute Black Top-Hat with
cdl.computation.image.morphology.compute_black_tophat()
- compute_erosion(param: MorphologyParam | None = None) None[source]
Compute Erosion with
cdl.computation.image.morphology.compute_erosion()
- compute_dilation(param: MorphologyParam | None = None) None[source]
Compute Dilation with
cdl.computation.image.morphology.compute_dilation()
- compute_opening(param: MorphologyParam | None = None) None[source]
Compute morphological opening with
cdl.computation.image.morphology.compute_opening()
- compute_closing(param: MorphologyParam | None = None) None[source]
Compute morphological closing with
cdl.computation.image.morphology.compute_closing()
- compute_all_morphology(param: MorphologyParam | None = None) None[source]
Compute all morphology filters using the following functions:
- compute_canny(param: CannyParam | None = None) None[source]
Compute Canny filter with
cdl.computation.image.edges.compute_canny()
- compute_roberts() None[source]
Compute Roberts filter with
cdl.computation.image.edges.compute_roberts()
- compute_prewitt() None[source]
Compute Prewitt filter with
cdl.computation.image.edges.compute_prewitt()
- compute_prewitt_h() None[source]
Compute Prewitt filter (horizontal) with
cdl.computation.image.edges.compute_prewitt_h()
- compute_prewitt_v() None[source]
Compute Prewitt filter (vertical) with
cdl.computation.image.edges.compute_prewitt_v()
- compute_sobel() None[source]
Compute Sobel filter with
cdl.computation.image.edges.compute_sobel()
- compute_sobel_h() None[source]
Compute Sobel filter (horizontal) with
cdl.computation.image.edges.compute_sobel_h()
- compute_sobel_v() None[source]
Compute Sobel filter (vertical) with
cdl.computation.image.edges.compute_sobel_v()
- compute_scharr() None[source]
Compute Scharr filter with
cdl.computation.image.edges.compute_scharr()
- compute_scharr_h() None[source]
Compute Scharr filter (horizontal) with
cdl.computation.image.edges.compute_scharr_h()
- compute_scharr_v() None[source]
Compute Scharr filter (vertical) with
cdl.computation.image.edges.compute_scharr_v()
- compute_farid() None[source]
Compute Farid filter with
cdl.computation.image.edges.compute_farid()
- compute_farid_h() None[source]
Compute Farid filter (horizontal) with
cdl.computation.image.edges.compute_farid_h()
- compute_farid_v() None[source]
Compute Farid filter (vertical) with
cdl.computation.image.edges.compute_farid_v()
- compute_laplace() None[source]
Compute Laplace filter with
cdl.computation.image.edges.compute_laplace()
- compute_stats() dict[str, ResultProperties][source]
Compute data statistics with
cdl.computation.image.compute_stats()
- compute_centroid() dict[str, ResultShape][source]
Compute image centroid with
cdl.computation.image.compute_centroid()
- compute_enclosing_circle() dict[str, ResultShape][source]
Compute minimum enclosing circle with
cdl.computation.image.compute_enclosing_circle()
- compute_peak_detection(param: Peak2DDetectionParam | None = None) dict[str, ResultShape][source]
Compute 2D peak detection with
cdl.computation.image.compute_peak_detection()
- compute_contour_shape(param: ContourShapeParam | None = None) dict[str, ResultShape][source]
Compute contour shape fit with
cdl.computation.image.detection.compute_contour_shape()
- compute_hough_circle_peaks(param: HoughCircleParam | None = None) dict[str, ResultShape][source]
Compute peak detection based on a circle Hough transform with
cdl.computation.image.compute_hough_circle_peaks()
- compute_blob_dog(param: BlobDOGParam | None = None) dict[str, ResultShape][source]
Compute blob detection using Difference of Gaussian method with
cdl.computation.image.detection.compute_blob_dog()
- compute_blob_doh(param: BlobDOHParam | None = None) dict[str, ResultShape][source]
Compute blob detection using Determinant of Hessian method with
cdl.computation.image.detection.compute_blob_doh()
- compute_blob_log(param: BlobLOGParam | None = None) dict[str, ResultShape][source]
Compute blob detection using Laplacian of Gaussian method with
cdl.computation.image.detection.compute_blob_log()
- compute_blob_opencv(param: BlobOpenCVParam | None = None) dict[str, ResultShape][source]
Compute blob detection using OpenCV with
cdl.computation.image.detection.compute_blob_opencv()