Processing Images#
This section describes the image processing features available in DataLab.
See also
Operations on Images for more information on operations that can be performed on images, or Analysis features on Images for information on analysis features on images.
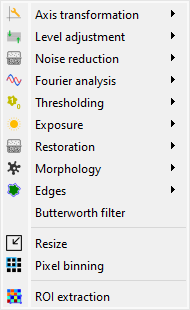
Screenshot of the “Processing” menu.#
When the “Image Panel” is selected, the menus and toolbars are updated to provide image-related actions.
The “Processing” menu allows you to perform various processing on the current image or group of images: it allows you to apply filters, to perform exposure correction, to perform denoising, to perform morphological operations, and so on.
Geometry#
Flip and rotation#
Create a new image by flipping or rotating the data of the selected image. The image may be flipped horizontally, vertically, or diagonally (transposition). It may be rotated by 90°, 270° or any user-defined value.
Operation |
Description |
|---|---|
Flip horizontally |
Mirror image along the vertical axis |
Flip diagonally |
Transpose the image (swap X/Y axes) |
Flip vertically |
Mirror image along the horizontal axis |
Rotate 90° right |
Rotate image 90° clockwise |
Rotate 90° left |
Rotate image 90° counter-clockwise |
Rotate by… |
Rotate image by any angle (user-defined) |
Distribute images along a grid#
Feature |
Description |
|---|---|
|
Distribute selected images on a regular grid |
|
Reset the positions of the selected images to first image (x0, y0) coordinates |
Axis transformation#
Set uniform coordinates#
Create a new image with uniform coordinates (i.e., with a constant pixel size in both X and Y directions).
Polynomial calibration#
Create a new image which is a polynomial calibration of each selected image with respect to Z axis.
Note
Linear calibration was removed from the UI as it is a special case of polynomial calibration with degree 1.
Level adjustment#
Normalize#
Create a new image which is the normalized version of each selected image by maximum, amplitude, sum, energy or RMS:
Normalization |
Equation |
|---|---|
Maximum |
\(z_{1} = \dfrac{z_{0}}{z_{\max}}\) |
Amplitude |
\(z_{1} = \dfrac{z_{0}}{z_{\max}-z_{\min}}\) |
Area |
\(z_{1} = \dfrac{z_{0}}{\sum_{i=0}^{N-1}{z_{i}}}\) |
Energy |
\(z_{1}= \dfrac{z_{0}}{\sqrt{\sum_{n=0}^{N}\left|z_{0}[n]\right|^2}}\) |
RMS |
\(z_{1}= \dfrac{z_{0}}{\sqrt{\dfrac{1}{N}\sum_{n=0}^{N}\left|z_{0}[n]\right|^2}}\) |
Clipping#
Apply the clipping to each selected image.
Offset correction#
Create a new image which is the result of offset correction on each selected image. This operation is performed by subtracting the image background value which is estimated by the mean value of a user-defined rectangular area.
Noise addition#
Generate new images by adding the same noise to each selected image. The available noise types are:
Noise |
Description |
|---|---|
Gaussian |
Normal distribution |
Uniform |
Uniform distribution |
Poisson |
Poisson distribution |
Noise reduction#
Create a new image which is the result of noise reduction on each selected image.
The following filters are available:
Filter |
Formula/implementation |
|---|---|
Gaussian filter |
|
Moving average |
|
Moving median |
|
Wiener filter |
Fourier analysis#
Zero padding#
Create a new image which is the result of zero padding on each selected image.
The following parameters are available:
Parameter |
Description |
|---|---|
Strategy |
Zero padding strategy (see below) |
Rows |
Number of rows to add (if strategy is ‘custom’) |
Columns |
Number of columns to add (if strategy is ‘custom’) |
Position |
Position of the added zeros: ‘bottom-right’, ‘centered’ |
Zero padding strategy refers to the method used to add zeros to the image, and it can be one of the following:
Strategy |
Description |
|---|---|
next_pow2 |
Next power of 2 (e.g. 512, 1024, …) |
multiple_of_64 |
Next multiple of 64 (e.g. 512, 576, …) |
custom |
Custom size (user-defined) |
Frequency-domain filters#
The following frequency-domain filters are available:
Method |
Description |
|---|---|
Butterworth |
Butterworth filter, based on skimage.filters.butterworth |
Gaussian filter |
Gaussian filter |
Thresholding#
Create a new image which is the result of thresholding on each selected image, eventually based on user-defined parameters (“Parametric thresholding”).
The following parameters are available when selecting “Parametric thresholding”:
Parameter |
Description |
|---|---|
Threshold method |
The thresholding method to use (see table below) |
Bins |
Number of bins for histogram calculation |
Value |
Threshold value |
Operation |
Operation to apply (> or <) |
The following thresholding methods are available:
Method |
Implementation |
|---|---|
Manual |
Manual thresholding (user-defined parameters) |
ISODATA |
|
Li |
|
Mean |
|
Minimum |
|
Otsu |
|
Triangle |
|
Yen |
Note
The “All thresholding methods” option allows to perform all thresholding methods on the same image. Combined with the “distribute on a grid” option, this allows to compare the different thresholding methods on the same image.
Exposure#
Create a new image which is the result of exposure correction on each selected image.
The following functions are available:
Function |
Implementation |
Comments |
|---|---|---|
Gamma correction |
||
Logarithmic correction |
||
Sigmoid correction |
||
Histogram equalization |
||
Adaptive histogram equalization |
Contrast Limited Adaptive Histogram Equalization (CLAHE) algorithm |
|
Intensity rescaling |
Stretch or shrink image intensity levels |
Restoration#
Create a new image which is the result of restoration on each selected image.
The following functions are available:
Function |
Implementation |
Comments |
|---|---|---|
Total variation denoising |
||
Bilateral filter denoising |
||
Wavelet denoising |
||
White Top-Hat denoising |
Denoise image by subtracting its white top hat transform |
Note
The “All denoising methods” option allows to perform all denoising methods on the same image. Combined with the “distribute on a grid” option, this allows to compare the different denoising methods on the same image.
Morphology#
Create a new image which is the result of morphological operations on each selected image, using a disk footprint.
The following functions are available:
Function |
Implementation |
|---|---|
White Top-Hat (disk) |
|
Black Top-Hat (disk) |
|
Erosion (disk) |
|
Dilation (disk) |
|
Opening (disk) |
|
Closing (disk) |
Note
The “All morphological operations” option allows to perform all morphological operations on the same image. Combined with the “distribute on a grid” option, this allows to compare the different morphological operations on the same image.
Edges#
Create a new image which is the result of edge filtering on each selected image.
The following functions are available:
Function |
Implementation |
|---|---|
Canny filter |
|
Farid filter |
|
Farid filter (horizontal) |
|
Farid filter (vertical) |
|
Laplace filter |
|
Prewitt filter |
|
Prewitt filter (horizontal) |
|
Prewitt filter (vertical) |
|
Roberts filter |
|
Scharr filter |
|
Scharr filter (horizontal) |
|
Scharr filter (vertical) |
|
Sobel filter |
|
Sobel filter (horizontal) |
|
Sobel filter (vertical) |
Note
The “All edges filters” option allows to perform all edge filtering algorithms on the same image. Combined with the “distribute on a grid” option, this allows to compare the different edge filters on the same image.
Erase area#
Erase an area in the image as defined by a region of interest (ROI).
Note
The region to erase is defined by the user through a dialog box. It can consist of a single ROI or multiple ROIs with various shapes (rectangular, circular, etc.). Note that the ROI defined here is not bound to any object; it is used solely to specify the area to erase in the image. In particular, it is independent of the image’s ROI, if any (the latter is shown in the dialog box as a masked area).
Resize#
Create a new image which is a resized version of each selected image.
Pixel binning#
Combine clusters of adjacent pixels, throughout the image, into single pixels. The result can be the sum, average, median, minimum, or maximum value of the cluster.
Resampling#
Generate new images by resampling each selected image. The following parameters are available:
Parameter |
Description |
|---|---|
math:x_{min} |
Minimum x-coordinate of the output image |
\(x_{max}\) |
Maximum x-coordinate of the output image |
\(y_{min}\) |
Minimum y-coordinate of the output image |
\(y_{max}\) |
Maximum y-coordinate of the output image |
Mode |
Image size definition mode: ‘Pixel size’ or ‘Output shape’. The ‘Pixel size’ mode allows to define the pixel size of the new image, while the ‘Output shape’ mode allows to define the number of pixels of the new image. |
ΔX |
Pixel size in x-direction (if ‘Pixel size’ mode is selected) |
ΔY |
Pixel size in y-direction (if ‘Pixel size’ mode is selected) |
Width |
Output image width in pixels (if ‘Output shape’ mode is selected) |
Height |
Output image height in pixels (if ‘Output shape’ mode is selected) |
Interpolation method |
Interpolation method to use: ‘nearest’, ‘linear’, ‘cubic’ |
Fill value |
Value to use for points outside the input image domain (if None, function uses NaN for extrapolation) |